By: Manepalli Ratna Sri, Surya Prakash and T Karuna
Normal human microbiota includes many fungi which are yeast and yeast-like [1]. These do not cause any disease under normal circumstances. However, they are opportunistic pathogens and can develop systemic and surface infections under some circumstances. The imbalance of microbiota composition in the body or irregularly functioning immune system are some of the main causes of fungal infection. Fungal infections can also be caused by long-term antibiotic treatment using broad spectral antibiotics, invasive medical procedures, long-term hospitalization, steroid treatment and [2].
Fungal infections generally occur in patients who are terminally ill and are immunocompromised. Due to this, the treatment has to be immediate and any delay in detection of the species of fungi causing infection may prove to be very fatal. Presently, the preliminary diagnosis is done through microscopic examination by human expert but it is very ambiguous as the species are visually very similar and can’t be differentiated easily. This is mainly due to existence of huge number of fungi patterns, minor variations in the patterns and their visual similarity, and human limitations in analysing such minute differences in the patterns.
To ascertain the type of fungi with greater confidence and to establish particular species they belong to, often additional biochemical tests, specialized molecular approaches and MALDI-TOF (Matrix Assisted Laser Desorption Ionization–Time of Flight) are recommended. This not only increases the cost of the diagnosis but also introduces the delay, sometimes up to 10 to 15 days. Further, biochemical tests have low sensitivity, so a pure isolate is required which increases the cost as well as delay. Molecular methods like FISH (Fluorescence in situ hybridization) and PCR (Polymerase chain reaction) are very expensive, species specific and require a lot of equipment. So, there is a need for a cost effective, accurate and efficient method for fungi species identification.
Recent developments in the field of deep learning have provided us the capability to deal with massive size of data collectively in a very efficient manner. Various deep learning architectures developed in last few years have made machines to perform even better as compared to humans in different application domains such as medical diagnosis etc. The developments in the field of deep learning have truly revolutionized the future of artificial intelligence. In recent years, deep learning has provided solutions to numerous difficult problems that has troubled the artificial intelligence community since long. Particularly, the ability of deep learning models to learn the hierarchical features from the images, makes it very powerful and capable of extracting high-level, complex characteristics as data representations. As mentioned in [3], deep convolutional networks are widely being utilized for analysis of medical images in many application areas like disease classification, computer aided diagnosis/retrieval, abnormality detection and segmentation.
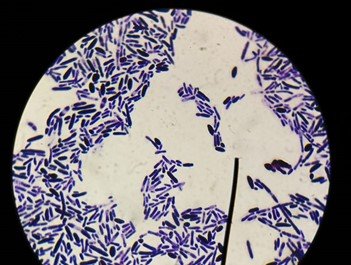
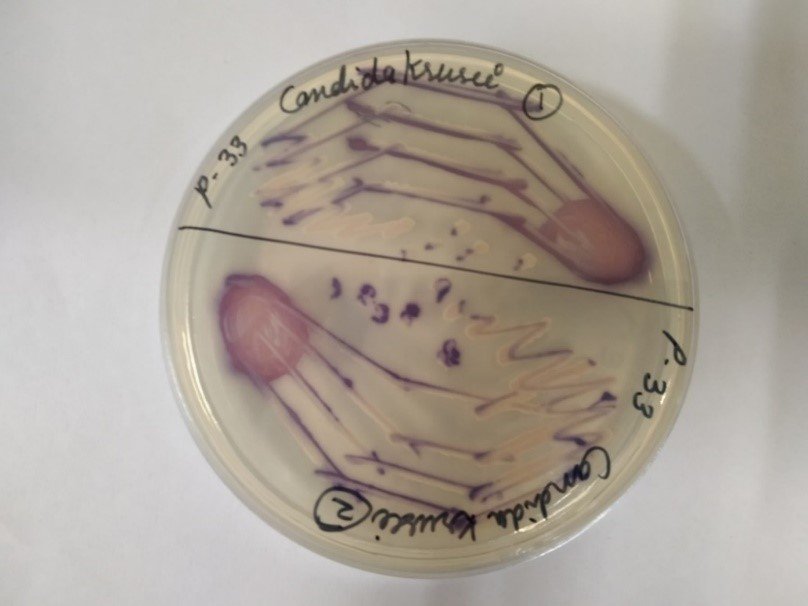
An AI tool, called FungalAITM [4], developed by an Australian group uses chest CT images. The group is more active with the surveillance and radiological approach to invasive fungal infections such as fungal pneumonia, but there is paucity of work with deep learning approach on the microscopic images of fungi which involves greater complexity due to similar patterns existing in fungal images. We can obtain two types fungal images namely microscopic and macroscopic images. Microscopic images are the images obtained when the fungi are seen under a microscope whereas macroscopic images are images obtained when seen through naked eye. Figures 1 and 2 show microscopic and macroscopic fungi images respectively.
In [5], authors propose an approach to perform segmentation of brain tumor utilizing Deep Neural Networks (DNN). They proposed a CNN architecture that simultaneously uses global as well as local contextual features. In [6], breast cancer classification is done through feed forward neural networks. This proposed method classifies breast cancer into benign and malignant. They have trained their network using the algorithm – island differential evolution propagation. In the work presented in [7], lung image patches with ILD (interstitial lung disease) is classified using a customized CNN with shallow convolution layer. The technique proposed in [8] accepts microscopic images and performs quantitative analysis of blood cells using CNN. The authors proposed a method to classify blood cells and do quantitative analysis using the YOLOv2 model. This approach is based on deep learning. The average accuracy of detection and segmentation of blood cells achieved is 80.6% and the precision achieved is 88.4%.
On similar lines, classification of fungi microscopic images can also be done leveraging the use of deep learning. Two approaches are proposed in [1] for classifying fungi microscopic images. The first approach is “fine-tuning the classifier’s block of the well-known network architectures”. It is based on a pre-trained DNN model. The DNN was initially pre-trained on instance classification. They fine-tuned the classifier’s block of some widely used network architectures like ResNet, InceptionV3, DenseNet169, and AlexNet. The networks were pretrained on the ImageNet Database. The second approach is using “Deep bag-of-words multi-step algorithm” consisting of image pre-processing, feature extraction using the convolutional layers of pre-trained DNNs, aggregation using Bag of Words, or Fisher Vector which is a more expressive modification of Bag of Words, and finally classification using Random Forest classifiers and support vector machine (SVM). Maximum accuracy achieved in [1] is 82.4%.
The pre-trained architecture models available cannot be used as fungal microscopic images are very different as compared to normal objects. Since they used the networks pre-trained on ImageNet, it did not give good results and they had to follow a long procedure involving aggregation and SVM for classification.
In [9] and in the review work presented in [10], automation of blood culture gram stain interpretation is proposed using Deep Convolutional Neural Network. They used Inception 3.0, which is a CNN previously trained to recognize everyday objects by Google. They retrained the classification layers only so that it can identify various standard Gram stain morphologies for positive smears of blood culture. The morphologies considered are chains of Gram-positive cocci, clusters of Gram-positive cocci, Gram-negative rods. Image Crops were formed and each crop was classified. Although this paper is not directly based on fungi image classification, we can extend the approach for classification of Fungi microscopic images with the required modifications. Also, image crops can be used to reduce the size of input image for feeding into the Neural Network.
In [11], another attempt is made for classification of few microbe species by utilizing some well-known DNN models. The images were cut into fragments and then fed to the DNNs. The database used had four species of fungi which belong to Candida genus – C. tropicalis, C. glabrata, C. dubliniensis, C. albicans which were stained using modified Gram method, resulting in 80 images in total. Good accuracy is obtained in the case of ResNet-50 when pre-trained (96%) and Xception when not pre-trained (95%). But, misclassifications occur in both the cases due to significant visual similarity in the microorganisms.
Open Research Problems:
Although a lot of work is being done in the medical image analysis field, fungi microscopic image classification is relatively a new area. The recent works show a lot of promise in this area but there are still some open challenges.
- Though the recent works show a good accuracy of classification, there is still scope to achieve improved classification accuracy. Accuracy is very important in the field of medical diagnosis and can’t be compromised. To replace the conventional methods with more efficient methods, accuracy needs to be the priority.
- There is a scale problem. The microorganisms have varied scales. Some are very small whereas some have a relatively larger size. So, it is difficult to fix a particular scale. Research has to be done to overcome this problem.
- Colour is also an important factor for identification of species and discoloration can give wrong results. There is an opportunity for research to overcome this problem.
- Currently, No computer database is available for fungi microscopic images although fungi atlas, textbooks, etc. are available. There is a requirement of a computer database of fungi microscopic images to facilitate further research.
- The popular pre-trained network architecture models available cannot be used as fungal microscopic images are very different from general object images. Research has to be done to develop a network architecture suitable for microscopic fungi images.
- There is a requirement for faster and accurate diagnosis of fungal infection which can be used by non-mycology experts in limited resources settings through tele-diagnosis mechanism. Hence, further research is required in this area.
References
[1] Zielinski, B., Sroka-Oleksiak, A., Rymarczyk, D., & Piekarczyk, A. (2019). Deep learning approach to describing and classifying fungi microscopic images. arXiv preprint arXiv:1906.09449.
[2] Rodrigues, C. F., Silva, S., & Henriques, M. (2014). Candida glabrata: a review of its features and resistance. European journal of clinical microbiology & infectious diseases, 33(5), 673-688.
[3] Anwar, S. M., Majid, M., Qayyum, A., Awais, M., Alnowami, M., & Khan, M. K. (2018). Medical image analysis using convolutional neural networks: a review. Journal of medical systems, 42(11), 1-13.
[4] “fungalAi| fungal diseases| machine learning | Australia”. Accessed on: 5 October 2021. [Online]. Available: https://www.fungalai.com/
[5] Havaei, M., Davy, A., Warde-Farley, D., Biard, A., Courville, A., Bengio, Y., … & Larochelle, H. (2017). Brain tumor segmentation with deep neural networks. Medical image analysis, 35, 18-31.
[6] Thein, H. T. T., & Tun, K. M. M. (2015). An approach for breast cancer diagnosis classification using neural network. Advanced Computing, 6(1), 1.
[7] Li, Q., Cai, W., Wang, X., Zhou, Y., Feng, D. D., & Chen, M. (2014, December). Medical image classification with convolutional neural network. In 2014 13th international conference on control automation robotics & vision (ICARCV) (pp. 844-848). IEEE.
[8] Tessema, A. W., Mohammed, M. A., Simegn, G. L., & Kwa, T. C. (2021). Quantitative analysis of blood cells from microscopic images using convolutional neural network. Medical & Biological Engineering & Computing, 59(1), 143-152.
[9] Smith, K. P., & Kirby, J. E. (2020). Image analysis and artificial intelligence in infectious disease diagnostics. Clinical Microbiology and Infection.
[10] Smith, K. P., Kang, A. D., & Kirby, J. E. (2018). Automated interpretation of blood culture gram stains by use of a deep convolutional neural network. Journal of Clinical Microbiology, 56(3), e01521-17.
[11] Zawadzki, P. (2020, September). Deep learning approach to the classification of selected fungi and bacteria. In 2020 IEEE 21st International Conference on Computational Problems of Electrical Engineering (CPEE) (pp. 1-4). IEEE.
Cite this article:
Manepalli Ratna Sri, Surya Prakash and T Karuna (2021) Classification of Fungi Microscopic Images – Leveraging the use of AI, Insights2Techinfo, pp.1
Also read:
- Humanoid Robots: The future of mankind
- Brain Computer Interaction (BCI): A Way to Interact with Brain Waves
- AI and Federated Machine Learning for Smart Healthcare